|
|
Post by kammy on Mar 20, 2010 14:40:08 GMT -5
Clathrin Coated Vesicles
Formed from Fibers While studying this Chlamydia .pdf: www.chlamydiae.com/docs/biology/biol_entry.aspit opened a lot of doors for me - we now have names for some of the artifacts we've been seeing and proves some of the theory that we have proposed is most likely happening, such as; we have HeLa cells, photos to show this aspect later.  Fig 2. A Chlamydia trachomatis elementary body (EB) in the process of entry into a HeLa cell. Tannic acid stained to enhance visualisation of chlathrin. Note the clearly demarcated outer envelope of the EB and the surrounding membrane (m) of the vacuole. Also note how small the clathrin coated pit (ccp) is compared to the chlamydial endosome. The bar represents 0.1 microns. We now have a name for our 'carbon-looking balls': The 'balls' that are inside of the fungus gnat larvae @100x: The same ones inside of our nematodes, Urine at 600x: They are called a clathrin coated pit, labeled "ccp" in the diagram above. If we research the clathrin coated pit, we see how certain fibers form these spheres:Reconstitution of Clathrin-coated Pit Budding from Plasma Membranes jcb.rupress.org/cgi/reprint/114/5/881.pdfThese photos show how certain fibers bend to form the circular vesicles: [/img][/center] A diagram from the NIH: www.ncbi.nlm.nih.gov/bookshelf/br.fcgi?book=mcb&part=A4895&rendertype=figure&id=A4895 |
|
|
|
Post by bannanny on Mar 20, 2010 15:17:08 GMT -5
Well kammy, you and jeany should be able to have your doctors there in Germany do the blood testing and skin biopsies to see if they match the frog's pathogens. I know I'd have a real hard time gettin whatever doc I end up with here to do it. So why don't you ask them? I know you guys will have a much better chance of getting certain tests done there in Germany than any of us will here in the US!
About the "carbon balls"... when I first started looking under a scope in the beginning of my disease those "balls" were the first thing I noticed under the scope. I kept telling anyone who wanted to look that it must have something to do with whatever I had. I hadn't come across this board yet or even found the morgellons site either. I was using a scope that wasn't digital back then too, so never got any pics of them... altho I've seen them numerous times in the pics I was finally able to take with my digital scope. Seems some of them form from the gel to me... or slime mold or whatever the gel is. But I know in my heart that whatever these metallic looking balls are, they're a huge part of morgs. I just know they are. I also saw tons of the black "rings" floating around everywhere back then... they seemed to form from the solid balls after awhile.
I saw alot of things back then that I'm still seeing today... which makes me angry cuz this thing should've been identified by now. If all of us can see these things, why can't the researchers? I actually think there's an unknown protein in this mess somewhere too that's responsible for alot of it.
hugs ~~ bannanny
|
|
|
|
Post by kammy on Mar 20, 2010 18:20:16 GMT -5
Here's the problem, Banny. For some reason it has been published that this Chytrid Fungus will not affect humans because it will not grow in above 31 degrees C. or 88.7 F. temperatures. I don't know who published this information but it has kept us with Morgellons away from looking at the frog's illness being related to ours. This is Morgellons Disease... anything is possible! Since the frog's dying is a big issue - there are many labs that will allow you to rub a swab across the tummy of your frog and mail the swab in for a PCR test. I'm going to swab my ear and send it in like my ear is a frog and see how the lab test comes back. There's a list of labs that will do this at the bottom of this article: www.amphibianark.org/chytrid.htmSince you don't have active lesions - it's possible just running the swab across your hands or feet might be able to give an indication, we just don't know at this time. |
|
|
|
Post by kammy on Mar 20, 2010 18:26:58 GMT -5
"Finding a Cure for Chytrid" www.sciencelearn.org.nz/Contexts/Saving-Reptiles-and-Amphibians/NZ-Research/Investigating-frog-disease"The frog research team at the University of Otago discovered that a common antibiotic (chloramphenicol) used for humans was able to kill samples of chytrid fungus in the lab. They decided to test this antibiotic on some Archey’s frogs that were infected with the disease. They found that they were able to cure the diseased frogs and keep them alive in the lab – an important breakthrough for this critically endangered species." Chloramphenicol en.wikipedia.org/wiki/Chloramphenicol"Chloramphenicol (INN) is a bacteriostatic antimicrobial. It is considered a prototypical broad-spectrum antibiotic, alongside the tetracyclines. Chloramphenicol is effective against a wide variety of Gram-positive and Gram-negative bacteria, including most anaerobic organisms. Due to resistance and safety concerns, it is no longer a first-line agent for any indication in developed nations, although it is sometimes used topically for eye infections; nevertheless, the global problem of advancing bacterial resistance to newer drugs has led to renewed interest in its use.[1] In low-income countries, chloramphenicol is still widely used because it is exceedingly inexpensive and readily available. Pharmacokinetics Chloramphenicol is extremely lipid soluble, it remains relatively unbound to protein and is a small molecule: it has a large apparent volume of distribution of 100 litres and penetrates effectively into all tissues of the body, including the brain. Chloramphenicol increases the absorption of iron.Mechanism of action Chloramphenicol is bacteriostatic (that is, it stops bacterial growth). It is a protein synthesis inhibitor, inhibiting peptidyl transferase activity of the bacterial ribosome, binding to A2451 and A2452 residues in the 23S rRNA of the 50S ribosomal subunit, preventing peptide bond formation. While chloramphenicol and the macrolide class of antibiotics both interact with ribosomes, chloramphenicol is not a macrolide. Chloramphenicol directly interferes with substrate binding, macrolides sterically block the progression of the growing peptide." www.answers.com/topic/chytridiomycosisTreatment options "It was reported in the June 8, 2009 issue of New Scientist that Reid Harris of James Madison University has found that coating frogs with Janthinobacterium lividum appears to protect them from chytridiomycosis.[10] Archey's frog Leiopelma archeyi, a critically endangered species endemic to New Zealand, was successfully cured of chytridiomycosis by applying chloramphenicol topically.[11]" |
|
|
|
Post by kammy on Mar 22, 2010 6:53:56 GMT -5
IDENTIFICATION and PROCESS BEHIND FORMATION OF THE MORGELLONS SPHERES FROM FIBERS These photos show how certain fibers bend to form the circular vesicles: [/img][/center] A diagram from the NIH: www.ncbi.nlm.nih.gov/bookshelf/br.fcgi?book=mcb&part=A4895&rendertype=figure&id=A4895This diagram from this website shows how the Morgellons fungi are most likely formed from our 'fibers': mycorhizes.com/Aarchaeosporales.htmlmycorhizes.com/images/classificationShema11.jpgresized to fit:  "Saccule sporifPre et spore d'Acaulospora rehmii Sporiferous saccule and spore of Acaulospora rehmii" or exhibit a spore dimorphism with the combination of acaulosporoVd and glomoVd types"   Spore of the glomoVd type (Glomus mosseae) "The organisms are characterise by a distinc molecular signature at the level of the SSU rRNA gene “YCTATCYKYCTGGTGAKRCG” corresponding to the homolog position 691 of the Jo1353 sequence of Saccharomyces cerevisiae and specific to the taxon. Families: Archaeosporaceae (Apendicispora, Archaeospora, Intraspora) and Geosiphonaceae (Geosiphon)" This is also the genus and species of the fungi involved in Morgellons, photo comparisons to follow. |
|
|
|
Post by jeany on Mar 22, 2010 8:56:43 GMT -5
  Main group: Glomeracecae - Glomeromycota belongs to family: Mykorrhiza  |
|
|
|
Post by jeany on Mar 22, 2010 9:14:22 GMT -5
Glomeromycota tolweb.org/Glomeromycota 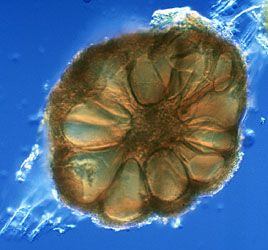 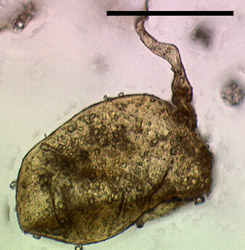 I've read on a German site (pdf file) that Glomeramycota extracts carbon from plant roots and was thinking that might be the 'metal looking' spheres, we've been seeing. Another possible aspect we should consider is, that this plant root fungus might have 'morphed or mutated' due to Agrobacterium which causes plant root galls and possibly 'created' a new type. The third pic looks def. morg-like to me. Jeany |
|
|
|
Post by kammy on Mar 22, 2010 9:19:37 GMT -5
Here is some information on the molecular sequences of the Chytrid fungi: www.biology.duke.edu/fungi/mycolab/publications/morehouseMolEcol2003.pdf"We sequenced 33 clones from the genomic library (average insert size ~ 950 bp). Polymerase chain reaction (PCR) primer pairs were designed to amplify eight of the cloned DNA regions using the software package Primer3 (Rozen & Skaletsky 1997). We focused primarily on clones that had significant matches to GenBank, under the assumption that coding regions were more likely to amplify single, homologous gene regions in contrast to noncoding regions that might be redundant or repetitive. Putative gene regions and P-values from blast searches were: cysteinyl tRNA synthase (ctsyn1;P= 3´10 - 13), anthranilate phosphoribosyltransferase (aprt13;P= 5´10 - 6), and 60S ribosomal protein (r6046;P= 6´10 - 13). Other targeted gene regions matched unidentified genes (bdc42;P= 10-12,uorf48;P=5´10 - 5) or did not show significant matches (bdc3, bdc33, rnap50). In addition, we examined sequences in GenBank to design primers to amplify translation elongation factor 1 a (tef1) and the nuclear subunit ribosomal RNA gene (lsu35). The sequences for the 33 clones have been deposited in GenBank (accession nos BH001009-BH001047). The following primers were used for PCR amplification (convention used is locus name followed by F and R designations for forward and reverse primers): ctsyn1 F (5 ¢ -ACCAACTATAACATCATCAAG-3 ¢), ctsyn1 R (5 ¢ -CGAATATCAGTCAACGCAAGC-3¢), aprt13 F (5 ¢ -GTCAGGGTTGGCTATTGTTCT-3 ¢), aprt13 R (5 ¢ -TGCTACTATTGCTGCTGTTGC-3 ¢), lsu35 F (5 ¢ -ATCCCTGTGGTAACTTTTCTG-3 ¢), lsu35 R (5 ¢ -ACGGACATGGGGAATCTGACT-3 ¢), r6046 F (5¢-CTATCTGCGCTCCCGTGTCAA-3¢), r6046R (5¢-AGGGCTGCAACAACTGGATTT-3¢), uorf48F (5¢-TCGAGGTGCAGACAAAACTTC-3¢), uorf48R (5¢-CAAACTGAGCCACAATAATGC-3¢), rnap50F (5¢-AATCCTATCCACCAGTTTCAG-3¢), rnap50R (5¢-TAACGATGAACGCCTTGTAGA- 3¢), bdc3F (5¢-TTCTGCTGCAAGAATCATCG- 3¢), bdc3R (5¢-AGTAGAAGCGGGTCGTTGAA-3¢), bdc33F (5¢-ATAGACCTTCGGGCTCTGGT-3¢), bdc33R (5¢-TTTCGTGTTAACCCAAAGGC-3¢), bdc42F (5¢- GGCCAACTTGTTGGATTTGT-3¢), bdc42R (5¢-TTGGAGCTCTGGTTCGACTT- 3¢), tef1F (5¢-TACAARTGYGGTGGTATYGACA-3¢), tef1R (5¢-ACNGACTTGACYTCAGTRGT-3¢)." |
|
|
|
Post by kammy on Mar 22, 2010 9:29:48 GMT -5
I'm still searching on how these 'beneficial' fungi are being used in forestry and agriculture methods. I believe they are putting them around tree roots...? mycorhizes.com/Asymbioses.html"In spite of a quite simple life cycle, those microscopic fungi live in symbioses with a majority of herbaceous plants and with several woody species. Arbuscular mycorrhizal symbioses profit to plant growth and plant protection against a variety of streses and as well to the propagation and the survival of microbial diversity, favoring the soil microbial equilibrium and consequently soil quality and sustainability through time." |
|
|
|
Post by jeany on Mar 22, 2010 9:44:17 GMT -5
How are mycorrhizal fungi re-established and how do you use
It's used as a 'fertilizer'.
Jeany
|
|
|
|
Post by kammy on Mar 22, 2010 9:46:49 GMT -5
Glomus tolweb.org/Glomeromycota Figure 4: Section of a sporocarp of Glomus sinuosum (isolate MD126, formerly Sclerocystis sinuosa). Spores are arranged around a center of interwoven hyphae and covered by a "peridium". Photo © Dirk Redecker, isolate courtesy of J. B. Morton at INVAM. Sporocarp diameter approximately 250 um. Here are two examples from a human specimen of a person recently diagnosed with PCV, Polycythemia Vera, this person smokes and is in poor health, also has COPD, at 100x, Arm Lesion cultured in nutrient agar. en.wikipedia.org/wiki/Polycythemia_vera(Not to say that PCV or COPD has anything to do with Morgellons.) I believe the Glomus sinuosum has mutated in this person: Glomus Stock Photo  |
|
|
|
Post by kammy on Mar 22, 2010 11:40:54 GMT -5
More Fiber to Spore Formation:
Diversispora, Pacispora mycorhizes.com/ADiversisporales.html[/img][/center] "The species belonging to this order are characterise by a distinct signature at the level of SSU rRNA gene “ YVRRYW/1-5/NGYYYGB” corresponding to the homolog position 658 of the JO1353 sequence of S. cerevisiae, 1346 of the JO1353 sequence “ GTYARDYHMHYY/2-4/GRADRKKYGWCRAC”, 650 of the JO1353 “TTATCGGTTRAATC” and 1481 of the JO1353 sequence “ACTGAGTTMATYT” specific to the taxon. Families: Diversisporaceae (Diversispora), Acaulosporaceae (Acaulospora and Kuklospora), Entrophosporaceae (Entrophospora), Gigasporaceae (Gigaspora and Scutellospora) and Pacisporaceae (Pacispora)." |
|
|
|
Post by kammy on Mar 22, 2010 11:49:17 GMT -5
Methods mycorhizes.com/Amethodologies.html"The methods used in the manipulation of arbuscular mycorrhizal fungi concerned the field sampling of soil samples from the rhizosphere, up to the fungal propagation, after the spore extraction from the substrate and the development of reference fungal collection. The practice of the proposed techniques require relatively few specialized equipment but needed some training in order to the practitionners to acquire enough dexterity to perform them, notably for the spore isolation, the in vitro cultivation and the microscopic observations. The briefly described methods herein presented are detailed in the following publications: Brundrett M, Bougher, N, Dell, B, Grove t, Malajczuk N 1996. Working with mycorrhizas in forestry and agriculture. ACIAR Monograph 32. 374 pages. Cranenbrouck S, Voets L, Bivort, Renard L, Strullu D.G., Declerck S 2005. Methodologies for in vitro cultivation of arbuscular mycorrhizal fungi with root organs. In: In vitro culture of mycorrhizas, Eds Declerck S, Strullu DG, Fortin A. Springer Verlag pages 342-375. Dalpé Y, Hamel C 2007. Arbuscular mycorrhizae Dans Manual of Soil Sampling and Methods of Analysis. 4rd Edition, Canadian Society of Soil Science Lewis Pub. of CRC Press. Chapter 30 pages 355-377. " |
|
|
|
Post by jeany on Mar 22, 2010 11:53:13 GMT -5
 I think it looks like our 'magnolia' we've been looking at....  |
|
|
|
Post by kammy on Mar 22, 2010 12:09:53 GMT -5
Identification Keys Last update: 2009 RETURN CODING AND ABBREVIATIONS used in the tables mycorhizes.com/Aidentification.htmlSpore colors: B = brown; BB = blackish brown BJ= brownish yellow CB = chocolate brown DB = dark brown DiB = dirty brown DY : dark yellow G : gray GB = golden brown GrB = gray brown H = hyalin O = orange OB = orange brown OR = orange red OY = orange yellow P = pink PB = pale brown PO: pale orange PY= pale yellow RB = reddish brown W = white Y= yellow Murograph: Am : amorph E : evanescent; F : flexible; L : laminate M : membranous muc : mucilaginous o ornamented * difficult to detect P : peridium; SF : semi-flexible SP : semi-permanent; U : unit X : extensible. Hyphal morphology c = cylindric co = constricted fl = flared fs= funnel-shape Pore: o = open p = close by a plug s = septate ? = unknown wt = wall thickened c = collar. KEYS KEY of Acaulosporaceae KEY of Archaeosporaceae KEY of Entrophosporaceae KEY of Glomeraceae KEY of Paraglomeraceae KEY of Ambisporaceae KEY of Diversisporaceae KEY of Gigasporaceae KEY of Pacisporaceae |
|
|
|
Post by kammy on Mar 22, 2010 12:14:01 GMT -5
Here's how the Chlamydia-like spore fits in there - mycorhizes.com/Aglossaire.htmlC Chlamydospores Thick walled spores ( from Greek Chlamys = envelope, surrounding) Emplacement photo 3 Thick walled spore of Glomus lamellosum We need to look at Chlamydospore and Glomus lamellosum! |
|
|
|
Post by kammy on Mar 22, 2010 12:40:18 GMT -5
|
|
|
|
Post by kammy on Mar 22, 2010 13:01:56 GMT -5
Look at the spore .gif file on the right move in when you open this page, www.agro.ar.szczecin.pl/~jblaszkowski/index.htmlyou can't copy it - it notes: swl1, swl2, swl3, & swl4 showing the cell layers with their names. I wonder if "swl..." is documented anywhere? I see this Oxford Journal .pdf that you have to pay to view, but here is a clue... [PDF] IN01-100: an allele of the Saccharomyces cerevisiae INO1 gene that ...Ino4, swl1, swl2 and swi3 mutants. Transcription of. INO1-100 was constitutive and independent of these regulators. A 20 bp deletion from -247 to -228 ... nar.oxfordjournals.org/cgi/reprint/23/8/1426.pdf That's why we're seeing the Morgellons pathogens in the red wine? It looks like they are possibly using this 'fiber to sphere' technology not only to create the fungi used in fertilizers but also in some of the yeasts used to ferment foods. S. cerevisiae now has several mutant versions, which are probably being used in wine, beer or bread making, this needs looked into - bread and beer need to be cultured. www.microbiologybytes.com/video/Scerevisiae.html"Yeasts such as Saccharomyces cerevisiae are single-celled fungi which that multiply by budding, or in some cases by division (fission), although some yeasts such as Candida albicans may grow as simple irregular filaments (mycelium). They may also reproduce sexually, forming asci which contain up to eight haploid ascospores. If you look closely at the video, you can see examples of budding cells (arrow, left). Saccharomyces cerevisiae has thick-walled, oval cells, around 10 µm long by 5 µm wide." Saccharomyces cerevisiae is commonly known as "bakers yeast" or "brewers yeast". The yeast ferments sugars present in the flour or added to the dough, giving off carbon dioxide (CO2) and alcohol (ethanol). The CO2 is trapped as tiny bubbles in the dough, which rises. Why does yeast do this? To gain energy from the breakdown (fermentation) of carbohydrates, as in the diagram opposite." |
|
|
|
Post by kammy on Mar 22, 2010 16:24:10 GMT -5
 I think it looks like our 'magnolia' we've been looking at....  This "Magnolia" is from a human lesion petri dish in which the dish had dried out... this shows a starving Glomus spore. |
|
|
|
Post by kammy on Mar 22, 2010 16:32:05 GMT -5
mycorhizes.com/A1993-2008.html1993-2008
The Molecular period "Since 1993, the development of molecular biology as a novel taxonomic tool allowed important break-through in the knowledge of arbuscular mycorrhizal fungi as in many other field of biology. Based on investigations made on the sequences of the SSU of the ribosomal DNA, a new phylum and new classe has been proposed, grouping together all the arbuscular mycorrhizal fungi into the Glomeromycota, within the class of Glomeromycetes, separating then definitively those fungi from the Zygomycotina, Zygomycetes. www.lrz-muenchen.de/~schuessler/amphylo/ The Glomeromycetes are then divided into 4 distinc orders, the Archeosporales, Diversisporales, Glomerales and the Paraglomerales. Those four orders are subsequently subdivided into 10 familiers including the Geosiphonaceae, a monogeneric family that include a unique species that result from the symbiosis between a blue-algae Nostoc and a Glomus species. The three families proposed by Morton and Benny (1990) are then found in the order Diversisporales (for Acaulosporaceae and Gigasporaceae) and Glomerales (for Glomus). The Glomeromycetes include actually 4 orders, 10 families and 13 genera."
|
|